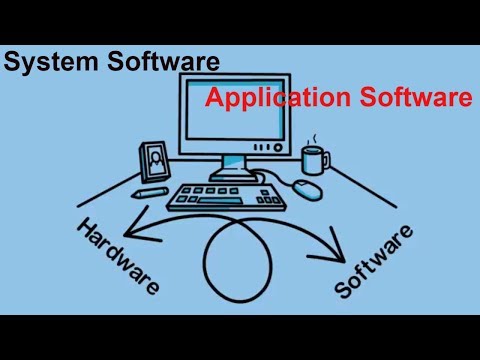

To learn more about how we are creating systems that streamline file management, economize processing power for complex files, and enable processing algorithms to work in the background to multitask getting to your omics results, read about our proteomics solution.This page is all about examples of bespoke software and how they are different compared to other software. 
Lastly, having one core location for all of the data reduces data security risks with single-point access control, audit trails, backup, and archival.All of the processing activities could be completed without interference from other users accessing from other locations or additional applications running in the background allowing for multi-tasked work.Connected processing applications like Proteome Discoverer could be automated to utilize files directly from the source location.Files wouldn’t need to be moved to be shared since users could access them for download through the browser interface.Starting with the core platform as a central file location, data from any connected acquisition application, such as Thermo Scientific Xcalibur, could directly save large omics files into the core and eliminate the need for a secondary network location altogether.I was happy to describe for the scientist how Thermo Fisher was rethinking and redesigning how software is structured from being one application that acquires, stores, secures, processes, and reports data, to a software structure with separate services and applications connected centrally. They had focused on reducing file risk and increasing the ability to share files with more people in the lab by centralizing to a network location to ensure that their users would not lose any critical information.īecause moving large raw files took so much time, what they were asking for was some kind of help to streamline the process, compress the files, or make the transfer easier (and faster). The scientist went on to describe a networking solution that they had put into place to allow file movement without external drive management. Then the results needed to be transferred into study summaries or removed to another location to make space on the processing computer. Files were acquired from instruments to associated acquisition PCs and those files had to be moved to the processing computer to generate results. To process proteomics data with Proteome Discoverer, the lab had set up a separate computer for the task. While I was minding the demonstration station, a scientist approached with a very specific and yet, identifiable omics data challenge that they were having with file management. 
As technology advances into detectors capable of taking wider swaths of readings at higher frequency, software is now tasked with managing terabytes of data produced in something like genomic sequencing over smaller file sizes produced by something like an IR spectrum. I was there to learn about and discuss data management solutions for the encroaching depth, breadth and size of data produced by a growing number of analytical instruments. Recently I attended the CHI hosted BIO-IT World conference in Boston.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
